|
You might also want to try the ‘Convoluted Background Subtraction’ tool available in the BioVoxxel Toolbox, which appears to do a better job at suppressing/removing the halos surrounding your stained cells, which potentially helps with the subsequent thresholding and particle selection.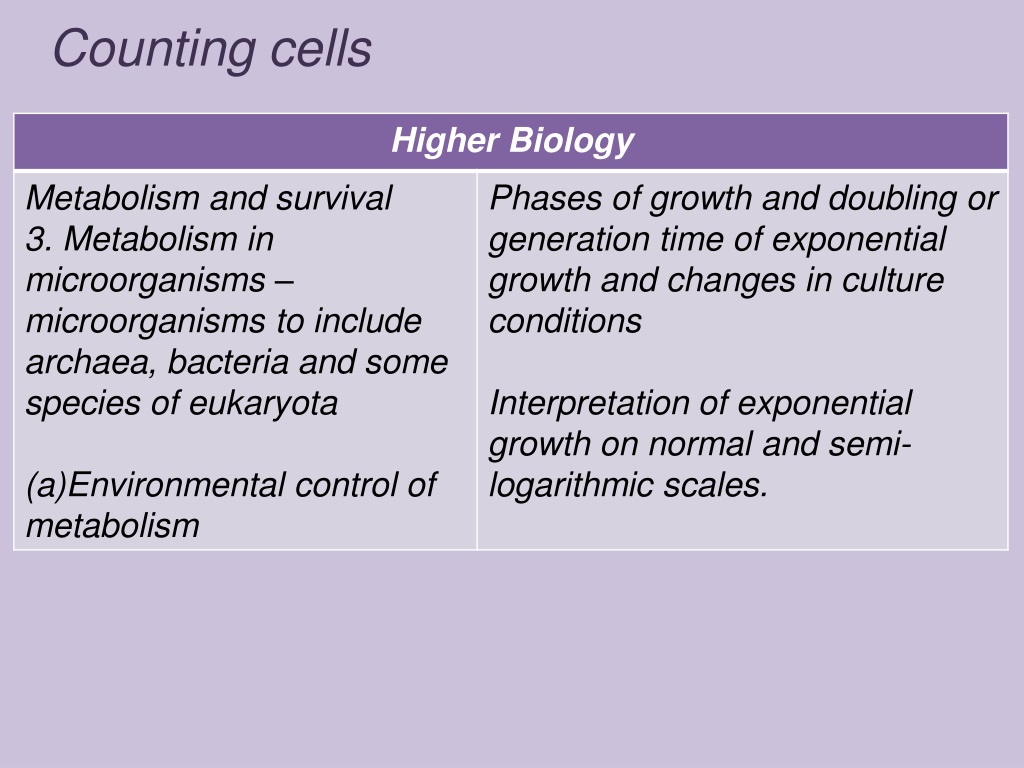 
Or you could smooth the particles on the thresholded image using binary filters to remove irregularities in the particles shape that cause the watershed algorithm to split single particles (I would expect single nuclei to be pretty smooth, round objects). So, the watershed step might not be required for your analysis.Īlternatively, you could try to use the ‘Irregular Watershed’ tool available in the Biovoxxel Toolbox ( BioVoxxel Toolbox). Looking at your sample image, the nuclei appear to be quite sparse with hardly any nuclei directly touching each other. When you say that you ‘can see lines within the cells’ that is due to the watershed and occurs if the structure has an irregular shape that is split by the watershed operation, which tries to separate touching particles. Screenshot of detected particles shown on original image: Run("Analyze Particles.", "size=5-150 circularity=0.65-1.00 display clear add") Volko run("Subtract Background.", "rolling=25") Either way, I don’t think that you will get any meaningful cell count from that clump and it is probably best excluded from the analysis. I assume that this is either some dirt or a larger clump of cells. The difficulty is the clump of material in the bottom right corner of the image. The code below hopefully gets you a bit closer to what you are aiming for. When you do the watershed, the options in the ‘Binary Options’ are also set such that the watershed is carried out on the black background rather than the white particles that you thresholded.įinally, your image doesn’t appear to be calibrated (scale set to 1 pixel = 1 inch). At the moment, you are selecting the halo around the centres of the stained structures rather than their centre - not sure whether this is your intention. How did you stain and image these nuclei?Īnyway, to threshold the bright spots that form the centre of your particles (I assume that this is what you try to count), I would suggest not to select the ‘Light Background’ option when you do the background subtraction. The stained particles also appear to have some rather strange halos more like a phase contrast image than a fluorescent image. Clicking “Create animaton” slows down the process further.Looking at your image, it looks a bit unusual for fluorescently labeled nuclei - I would expect the contrast of the nuclei relativeto the background to be considerably higher for most of the conventional fluorescent nuclei stains (e.g. The analysis will take very long if the Max value is 255 and over. If too many or not enough particles were counted, adjust the Max value and/or radius. Usage: Open the Cell Counter plugin and the image/stack you want to count (if the Cell Counter plugin is already open you dont need to open a new instance). Tick “Show progression messages” and press “Start Watershed”. Select “Overlaid dams” from the bottom menu: This accounts for the fact that ellipsoid shapes which will be counted can be touching but will still be included in the analysis (a function which the regular particle counter plugin in ImageJ is lacking): Select “ 8-connected” particles for the analysis. If too many cells/particles are counted, reduce the max down to 175. If the staining is faint and not enough cells are picked out, in particular for the hypertrophic zone, increase max to 210-220. 
0 to 200 is a good setting for the general DAPI staining. Min/max level indicates the intensity of greyness which is still recognised as a particle. It works well at 0.5 for the resting and proliferative zones and 1.0 for the hypertrophic zone (larger cells). Radius indicates the predicted radius of the particles to be measured. 
There are certain parameters which need to be set before the analysis: To quantify DAPI positive cells open the file and select the zone of interest:Ĭlear the outline leaving only the zone of interest:įor DAPI and other fluorescence images, invert the colours:Īdjust brightness and contrast (clicking “Auto” and then “Apply” is often enough): Make sure java is up to date on the computer as this is a java applet. Watershed algorithm can be used both in ImageJ or Fiji. Images should be taken at the right magnification, allowing identification of individual cells in each zone of the growth plate.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
